|
|
Post by skyship on Mar 28, 2010 17:29:51 GMT -5
Abstract A method for producing a recombinant baculovirus expression vector , capable of expression of a selected gene in a host insect cell, is disclosed. The preferred method involves cleaving baculovirus DNA to produce a DNA fragment comprising a polyhedrin gene or portion thereof, including a polyhedrin promoter. A recombinant shuttle vector is prepared by inserting said DNA fragment into a cloning vehicle and thereafter inserting a selected gene into the thus modified cloning vehicle such that it is under the transcriptional control of the polyhedrin promoter. The recombinant shuttle vector is contacted with a baculovirus DNA so as to effect recombination and incorporation of the selected gene into the baculovirus genome. www.patentstorm.us/patents/4745051.htmlpolyhedron promoter: In view of this, the promoter has been used to construct expression vectors to allow high level of expression of any gene under the influence of this promoter. This is being used for developing biopesticides or even for production of specific chemicals in industry. www.molecular-plant-biotechnology.info/cloning-and-expression-vectors/polyhedrin-promoter-from-baculovirus.htmskyship |
|
|
|
Post by skyship on Mar 28, 2010 17:46:50 GMT -5
polyhedron gene: silk worm fibroin: Studies on Advanced Fiber/Textile Science and Technology New Biological Fiber Alteration of Silkworm Fibroin using Gene Targeting Accession number;04A0711079 Title;Studies on Advanced Fiber/Textile Science and Technology New Biological Fiber Alteration of Silkworm Fibroin using Gene Targeting Author;NAKAGAKI MASAO(Shinshu Univ., Textile Sci. and Technol., JPN) TAKEI RYUZO(Shinshu Univ., Textile Sci. and Technol., JPN) KIGUCHI KENJI(Shinshu Univ., Textile Sci. and Technol., JPN) KAJIURA ZENTA(Shinshu Univ., Textile Sci. and Technol., JPN) YAJIMA MASAO(Shinshu Univ., Textile Sci. and Technol., JPN) Journal Title;Studies on Advanced Fiber/ Textile Science and Technology. Grant-in-Aid for COE Research Journal Code:N20041669 ISSN: VOL.;NO.;PAGE.101-102(1999) Figure&Table&Reference; Pub. Country;Japan Language;Japanese Abstract;The strong promotor of the polyhedron gene of baculovirus (NPV) was used to remodel fibroin gene, and the recombinant NPV that has base sequence of fibroin gene of the silkworm chromosome before and after the foreign gene was prepared. After making the recombinant NPV carry the foreign gene to the nucleus of reproductive cells, the foreign gene was inserted into the fibroin gene using homologous recombination between the fibroin gene sequence on viral DNA and the fibroin gene on chromosomes. The study also involved the examination of the strong promotor of polyhedron gene of NPV. sciencelinks.jp/j-east/article/200423/000020042304A0711079.php=================== polyhedron gene and human infections?       ? Here is the problem with this particular construction: =============================== Abstract Three recombinant baculoviruses which are capable of expressing human immunodeficiency virus (HIV) protease, p55gag, and both products simultaneously in insect cell culture have been constructed. Upon co-infection of cells with the protease and p55gag-expressing viruses, authentic processing of the gag precursor is observed to take place. This processing could be reproduced in vitro using mixtures of cellular lysates containing the expressed proteins. When expressed alone, uncleaved p55gag precursor appears to form retroviral core-like particles within the cytoplasm of infected cells. Metabolic labeling studies of the baculovirus-expressed gag products have demonstrated that p17 is myristylated at its amino terminus, and that p24 is phosphorylated. In these respects, the insect cell system is evidently capable of carrying out post-translational processing resembling that which occurs in authentic HIV-1 replication. tinyurl.com/yb3cgyf======================================= insects could carry HIV. would be vectors of the HIV. However, we do not test positive for HIV. ============================== Use of baculovirus early promoters for expression of foreign genes in stably transformed insect cells United States Patent 5077214 www.freepatentsonline.com/5077214.htmlThe present invention provides an alternative strategy for baculovirus-mediated foreign gene expression: the promoters from immediate-early or delayed-early baculovirus viral genes were used to obtain the continuous expression of foreign genes in stably-transformed insect cells. The IE1 or the 30K promoters from Autographa californica nuclear polyhedrosis virus can drive the continuous expression of a foreign gene in insect cells. IE1-β-galactosidase or 39K-β-galactosidase constructs were used in combination with the IE1-neomycin resistan The Government may have rights in this invention pursuant to a funding agreement with the National Science Foundation (NSF), Grant No. DMB-88 04732. skyship |
|
|
|
Post by skyship on Mar 28, 2010 18:03:25 GMT -5
Now could there be other use of baculovirus that would infect humans other than HIV? ONe way is hepatocytes, liver in humans. However this is used for gene therapy. ============================= Note the abstract for quick view. www.pnas.org/content/92/22/10099.full.pdf-======================== The is what is involved: Baculovirus Autographa californica nuclear polyhedrosis virus: or named (AcNPV) but used for gene therapy, viruses are used to mutate the correct gene in in gene therapy in diseaeses of liver, but, that now is being changed to prevent the viral component from staying in the body. so, far baculo does not directly infect humans. ====================== Health Risks, this explains................ more........... ....." Recombinant baculovirus containing Bacillus thuringiensis toxin have not proven successful in controlling insect pests (Martens et al 1995). However, recombinant baculovirus modifying juvenile hormone proved effective in insect control (Bonning et al 1999). Recombinant baculovirus containing an antisense fragment to the c-myc oncogene proved effective in target insect control (Lee et al 1997). The behavior of the myc oncogene recombinant vector bears careful study regarding non-target animals and its impact during human lliver infection. Baculovirus vectors efficiently transfer genes into human liver cells (Hofmann et al 1995; Boyce and Bucher 1996). The vectors transferred into human liver tissues most effectively in perfused liver tissue because serum components hampered virus transfer (Sandig et al 1996).Human conditions causing defects in complement should allow liver transfer of recombinant baculovirus. Inhibitors of complement facilitate baculovirus gene transfer (Hofmann and Strauss 1998). Hybrid baculovirus-adeno virus vectors have been used to deliver genes to human cells (Palombo et al 1998). Baculovirus vectors have been used to deliver hepatitis B to human liver efficiently to allow study of hepatitis B drug therapy (Delaney et al 1999). In conclusion baculovirus vectors are being used to control insect pests because they are effective and persist for a long time in the environment. Baculovirus vectors are also being used in gene therapy of human liver. These areas of research seem to exist as two solitudes and the risks of one are not evaluated in the context of the other. The most disconcerting finding is the one showing that replication of the baculovirus is inherently unpredictable. However, there may be some who believe that we should all have unlabelled liver gene therapy with our salad. "..... www.biotech-info.net/gene_therapy.htmlskyship www.biotech-info.net/gene_therapy.html |
|
|
|
Post by skyship on Mar 28, 2010 18:15:18 GMT -5
Other ways baculovirus could infect humans?    ?? "Baculovirus is a circular DNA duplex, it replicates in the insect cell nucleus and replication is prone to the generation of defective genomes by deletion. The mode of virus replication seems to make the recombinant virus highly unpredictable and prone to generating potentially undesirable variants. This important finding has not yet influenced the risk analysis of recombinant baculovirus insecticides and gene therapy vectors." However, this again could be tested for. Morgellons can't ....... Because it involves our receptors taking in folding and unfolding through prions. Prions are the vectors, a created vector. By this I mean: Sup35p which has the potential to form amyloids, tau.......not just in brain, but in every area of body. skyship |
|
|
|
Post by skyship on Mar 28, 2010 18:17:42 GMT -5
Here is a paper I wrote: Yeast prion as Operon in Morgellons organisms.
by
(Skyship) Prions are the non-genetic carriers and implementors of DNA modification, genetic engineering, genetic control through forced or directed evolution in the environment. These are foreign adaptation instruments for reengineering human genes. It appears that a reengineering project has been implemented to introduce artificial transformation of the human genome. Due to many differences in the human species, a universal plan has been constructed to implement engineered DNA through the release of foreign adaptive protein constructions into the environment, foods, and humans. The prion which becomes a self replicating machine, is being used as a tool to implement gene alterations and to integrate self assembling organisms into human bodies, replacing native genes and/for use in controlling the transformed human. Prion proteins are found in such organisms as Yeast, Podospora anserina and Aplysia Californica. The yeast used is sup35p among others. Sup35p is described here: "Amyloid formation of a yeast prion determinant. T Scheibel The [PSI+] factor of the yeast Saccharomyces cerevisiae is a cytoplasmic, epigenetic regulator of translation termination and can be transmitted from mother to daughter cells in a non-Mendelian manner. The transmission is caused by self-perpetuating noncovalent changes in the physical state of the protein determinant Sup35p, rather than by changes in its encoding gene. This phenomenon is reminiscent of the protein-only mechanism proposed for the infectious agent in a group of unusual, fatal neurodegenerative diseases in mammals. These diseases, known as prion diseases, are thought to involve a self-perpetuating change in the conformation of the prion protein (PrP) from a soluble form to one reflecting amyloid structure. In contrast to mammalian PrPs, Sup35p[PSI+] is not associated with disease in yeast and is not infectious for humans. Because of the mechanistic similarities of transmission of a specific, nonsoluble protein conformation, the epigenetic inheritance of [PSI+] in yeast was called a yeast prion phenomenon, and the yeast prion hypothesis was born. The elucidation of the mechanism by which alternative protein conformations transmit their structural information is key to understanding how proteins function as elements of epigenetic inheritance and how amyloidogenic conformations can be propagated. Yeast provides an ideal system to analyze both the epigenetic traits in vivo and the phenomenon of amyloid formation in vitro. The combination of these tools will help to determine the general mechanism of prion and amyloid appearance and propagation." www.epidna.com/showabstract.php?pmid=15126688However, these are just as infectious as the prp(sc), and have the power to integrate into human proteins as shown by this report of 6 infectious prions which change chromatin remodeling. In otherwords, this changes our chromatin which are inside the gene structure. The prion itself does not make the gene change, but carries the gene altering proteins and enzymes which are introduced into the unfolded prion we already have, the prp(c). It is the prion that inserts itself with its DNA or RNA package and replicates using the new RNA or DNA transference. "The yeast global transcriptional co-repressor protein Cyc8 can propagate as a prion. Patel BK, Gavin-Smyth J, Liebman SW. Laboratory for Molecular Biology, Department of Biological Sciences, University of Illinois at Chicago, 900 S. Ashland Avenue, Chicago, Illinois 60607, USA. * Nat Cell Biol. 2009 Mar;11(3):241-3. Although many proteins can misfold into a self-seeding amyloid-like conformation, only six are known to be infectious, that is prions. The prions [PSI(+)], [PIN(+)], [URE3], [SWI(+)] and [HET-s] cause distinct heritable physiological changes in fungi, whereas PrP(Sc) causes infectious encephalopathies in mammals. It is unknown whether 'protein-only' inheritance is limited to these exceptional cases or whether it represents a widespread mechanism of epigenetic control. Towards this goal, we now describe a new prion formed by the Cyc8 (Ssn6) protein of Saccharomyces cerevisiae. Analogously to other yeast prions, transient overproduction of a glutamine-rich region of Cyc8 induced a heritable dominant cyc8(-) phenotype that is transmitted cytoplasmically and is dependent on the chaperone Hsp104 and the continued presence of the Cyc8 protein. The evolutionarily conserved Cyc8-Tup1 global transcriptional repressor complex forms one of the largest gene regulatory circuits, controlling the expression of more than 7% of yeast genes. Our finding that Cyc8 can propagate as a prion, together with a recent report that Swi1 of the Swi-Snf global transcriptional regulatory complex also has a prion form, shows that prionization can lead to mass activation or repression of yeast genes and is suggestive of a link between the epigenetic phenomena of chromatin remodelling and prion formation." tinyurl.com/yf6qxqzwww.ncbi.nlm.nih.gov/pubmed/19219034?ordinalpos=1&itool=EntrezSystem2.PEntrez.Pubmed.Pubmed_ResultsPanel.Pubmed_SingleItemSupl.Pubmed_Discovery_RA&linkpos=2&log$=relatedarticles&logdbfrom=pubmed Prion proteins are found in such organisms as Sacromycetes Ceraviseae, Podospora anserina and aplysia californica. Other organisms also carry the prion replicating machine. The yeast prion is the determinant. " The elucidation of the mechanism by which alternative protein conformations transmit their structural information is key to understanding how proteins function as elements of epigenetic inheritance and how amyloidogenic conformations can be propagated. Yeast provides an ideal system to analyze both the epigenetic traits in vivo and the phenomenon of amyloid formation in vitro. The combination of these tools will help to determine the general mechanism of prion and amyloid appearance and propagation." www.epidna.com/showabstract.php?pmid=15126688 This can then be used for the introduction of molecular tools for reengineering. The tools become the conveyor for the transformation of cells at a smaller level. This involves changing genes from the inside out. However, genes condense in the chromatin causing bundles to form. In Morgellon's Disease, fibrous bundles are present in skin. Could this be the chromatin bundles formed by the prion replicating foreign adaptive proteins from yeast, fungi, and slug prions? It appears as if the bundles leaving the body through the skin are being rejected. Are the created bundles being rejected or are the native bundles being replaced by the artificial fibrous bundles of altered chromatin? To prove that prions are the catalyst for altering, modifying and exchanging RNA or DNA in native human protein for artificial nuclei in cells by way of using proteins and recombinant transferance may be a hard one to prove. However, ignoring the fact that prions of any sort, both bse, tse, and the above formed from proteins used for replication in unfolded prion proteins is a way of altering the very structure we know as human. The foreign adaptive organism that causes Morgellon's Disease is constructed from biological elements, chemical elements, geological elements, and elements of physics. To bring forth first, the biological elements one has to consider the properties and the characteristics of prions and the probability of creating an organism that operates like a machine and can then be replicated with polymers, metals, and nanotubes. The models for gene exchange, horizontal transfer, lateral transfer and genetic mobile elements found in spores, and metal particles in aerosol operations can initiate the prion operations. This is where the filament of life meets the spark of life, through the cell and the nucleosus or center of the cell and its mitochondrial formation. It appears the yeast prion and its amyloid forming abilities are at the core of this phenomena we call Morgellons. In later papers, we will discuss how other proteins from nature, from chemical reactions, from geological, mineral, metal, and physics positive and negative electrons cause the immune system to go down. Mycoplasma in it's many forms affects the energy of humans. references: www.epidna.com/showabstract.php?pmid=15126688www.pitt.edu/~pvdwel/prot_amyloid.htmlwww.ncbi.nlm.nih.gov/pubmed/7909170?dopt=Abstractwww.physorg.com/news173974213.htmlwww.epidna.com/showabstract.php?pmid=11931236www.google.com/search?ie=UTF-8&oe=utf-8&q=epigenetic+phenomena+of+chromotin+remodeling+of+prion+formationwww.ncbi.nlm.nih.gov/pubmed/19219034?ordinalpos=1&itool=EntrezSystem2.PEntrez.Pubmed.Pubmed_ResultsPanel.Pubmed_SingleItemSupl.Pubmed_Discovery_RA&linkpos=2&log$=relatedarticles&logdbfrom=pubmed Skyship |
|
|
|
Post by skyship on Mar 28, 2010 18:47:54 GMT -5
Proteins acting as genes: Prions of yeast as epigenetic phenomena: high protein "copy number" inducing protein "sile "Yeast infectious protein (prion) forms of the Ure2 and Sup35 proteins determine the nonchromosomal genes [URE3] and [PSI], and these are, therefore, the basis for a kind of epigenetic phenomena. In many systems, introduction of multiple copies of a DNA gene, or dsRNA copies of its sequence, results in the epigenetic silencing of that gene. In parallel with these homology effects, which act at the level of DNA or RNA, elevated copy number of the Ure2 and Sup35 proteins increases the frequency of their own "silencing" by prion formation. Both [URE3] and [PSI] appear to be due to self-propagating-amyloid formation of Ure2p and Sup35p, respectively. Another prion, [Het-s] of the filamentous fungus, Podospora anserina, is necessary for a normal cellular function, heterokaryon incompatibility. Since these prions are nonchromosomal genes, they are proteins acting as genes, a parallel to the fact that nucleic acids can catalyze enzymatic reactions." www.epidna.com/showabstract.php?pmid=11931236skyship |
|
|
|
Post by skyship on Mar 28, 2010 19:19:29 GMT -5
Kammy, I do find this tree of life very interesting: It has Baculovirus at the bottom here: Here it has it's own little niche: =========================== Fig 18: Evolutionary tree of DNA polymerase amino termini (Villareal and Defilippis) (click to enlarge). Villareal and Defilippis have likewise investigated the idea that DNA viruses are the origin of DNA replication proteins, by investigating the amino terminus and constructing an evolutionary tree which shows DNA polymerases of viruses Eukaryotes, archaea and some bacteria and phages rooted in a tree consistent with a viral origin. This idea has a great deal of plausibility because viruses are now know to have a potentially primal origin, rather than being recent escapees from cellular genomes which have undergone reductive parasitic changes to their genome. Viruses clearly also have retained both RNA-RNA, DNA-DNA and retrotranscription DNA-RNA-DNA using both RNA and DNA stages in their capsid viral forms, so they retain all the transitional states between RNA and DNA-based replication. www.dhushara.com/book/unraveltree/DNApolamino.png============================== another tree here: ====================== Fig 19: Bacterial DNA polymerases also show viral members (underlined) close to the root of the tree (File et. al.). Furthermore the retroviruses and related mobile genetic elements have a common ancient evolutionary origin, which is related to telomerase, which itself uses an RNA primer to initiate chromosome duplication. There is thus a plausible case that telomerase is in fact a biological fossil of a retroviral conversion of the founding Eukaryote cell line to a DNA genome. www.dhushara.com/book/unraveltree/unravel.htmskyship |
|
|
|
Post by katinka on Mar 28, 2010 20:05:10 GMT -5
Very good paper you wrote there, Sky! YES! Kammy and I were looking at Saccharomyces cerevisiae after we cultured red wine samples. According to previous research I found it most interesting that Saccharomyces cerevisiae promotes actin development which is a protein involved in the production of Amyloids and that two proteins Sup35 and Hsp104 of this specific yeast has the ability to 'create' an overproduction of Amyloids in humans causing diseases such as Alzheimer's, related to prions, for example. I wrote a blog entry on amyloids when I was looking at Falpe and p10 proteins after we 'discovered' the Baculovirus and how it has the capability to from cytoplasmic fibers in insects and also infect humans causing a form of cytoplasmic fibrils namely amyloids. morgellons2.wordpress.com/2009/09/12/formation-of-actin-filaments-in-mammalian-cells-baculovirus-protein-falpe-and-p10/Here is a picture of sup35 fibers: 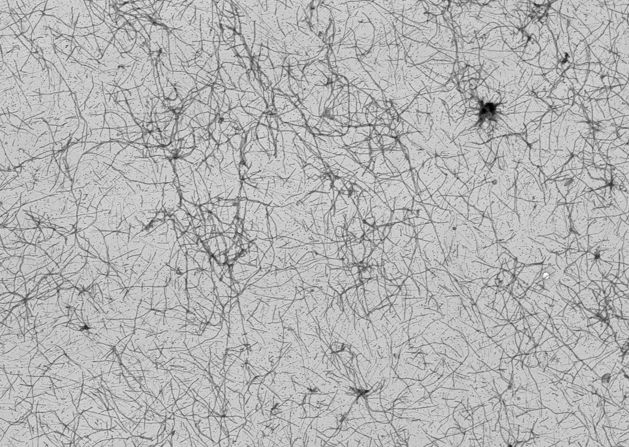 If you don't mind, I would like to add our previous information on this subject here: Quote from LB: According to my previous research on Amyloid fibrils I was thinking about this unknown protein Carnicom mentioned. I have found that a yeast called Saccharomyces cerevisiae promotes actin development which is a protein involved in the production of Amyloids. I found interesting that two proteins Sup35 and Hsp104 of this specific yeast has the ability to 'create' an overproduction of Amyloids in humans causing diseases such as Alzheimer's for example. If you look at the picture Kammy posted above the Amyloids do look similar to Morgellons fibers. I personally believe that the Morgellons fibers are a body waste product resulting from this mentioned overproduction of Amyloids. Researchers Discover Protein That Dissolves Amyloid Fibers www.sciencedaily.com/releases/2004/05/040521070408.htmThe finding follows years of study that has focused on a yeast protein called Sup35, a protein that helps cells translate genetic information into strings of amino acids – the building blocks of protein molecules. Sometimes Sup35 suddenly forms amyloid fibers similar to those found in Alzheimer's patients. In yeast, however, this doesn't kill the cell. Rather, it is part of the cell's normal biology, changing the types of proteins that the cell makes. Previous research in the Lindquist lab described how a protein called Hsp104 seemed to affect Sup35's ability to form amyloid fibers. When a yeast cell contained either high amounts of Hsp104 or none at all, amyloid fibers never formed. But when Hsp104 levels were small, the fibers flourished. In this new study, Lindquist and postdoctoral researcher James Shorter isolated Sup35 and Hsp104. Here they saw that small amounts of Hsp104 catalyzed the formation of amyloid fibers, but large levels of the protein actually caused the fibers to dissolve. Prions, those infectious proteins implicated in conditions such as mad cow disease, are a subclass of amyloids. In yeast cells, Sup35 technically is a prion. Hsp104 belongs to a class of proteins that sometimes are influenced by environmental factors. A yeast cell in one type of environment can experience an abundance of Hsp104, which would then keep Sup35 from forming amyloid fibers in that cell. But put that cell in a different environment and the result may be a more moderate level of Hsp104 that would, in turn, create amyloid fibers in Sup35, changing how that protein functions and ultimately altering the cell's biology. Saccharomyces cerevisiae en.wikipedia.org/wiki/Saccharomyces_cerevisiaeIt is believed that it was originally isolated from the skins of grapes (one can see the yeast as a component of the thin white film on the skins of some dark-colored fruits such as plums; it exists among the waxes of the cuticle). It is one of the most intensively studied eukaryotic model organisms in molecular and cell biology, much like Escherichia coli as the model prokaryote. It is the microorganism behind the most common type of fermentation. Saccharomyces" derives from Latinized Greek and means "sugar mold" or "sugar fungus", saccharo- being the combining form "sugar-" and myces being "fungus", Killer yeasts en.wikipedia.org/wiki/Killer_yeastsKiller yeasts are yeasts, such as Saccharomyces cerevisiae, which carry a double-stranded RNA virus causing them to secrete a number of toxic proteins which are lethal to receptive cells. These yeast cells are immune to the toxic effects of the protein due to an intrinsic immunity, The virus, L-A, is an icosahedral virus of S. cerevisiae comprising a 4.6 kb genomic segment and several satellite double-stranded RNA sequences, called M dsRNAs. The genomic segment encodes for the viral coat protein and a protein which replicates the viral genomes.[4] The M dsRNAs encode the toxin, of which there are at least three variants in S. cerevisiae, The two most studied variant toxins in S. cerevisiae are K1 and K28. K1 binds to the β-1,6-D-glucan receptor on the target cell wall, moves inside, and then binds to the plasma membrane receptor Kre1p. It forms a cation-selective ion channel in the membrane, which is lethal to the cell. K28 uses the α-1,6-mannoprotein receptor to enter the cell, and utilizes the secretory pathway in reverse by displaying the endoplasmic reticulum HDEL signal. From the ER, K28 moves into the cytoplasm and shuts down DNA synthesis in the nucleus, triggering apoptosis. Young and Yagiu (1978) experimented with methods of curing killer yeasts. They found that using a cycloheximine solution at 0.05 ppm was effective in eliminating killer activity in one strain of S. cerevisiae. Incubating the yeast at 37°C eliminated activity in another strain. Many toxins are sensitive to pH levels; for example K1 is permanently inactivated at pH levels over 6.5. ** ph buffering?? How I came to my latest conclusions: I am studying this sup35 fiber and seeing that it is being called a 'release factor', which is exactly the way I have described it. It is fungus-like (Chlamydia-like?)... it branches out and spreads to form the carbon capsids. www.med.upenn.edu/shorterlab/Papers/SweenyPRION.pdf''Infectious amyloid forms of the release factor, Sup35, comprise the yeast prion [PSI+].'' A yeast prion called PSI+ might be involved?... I do a search on PSI+ in which I look at Wikipedia first because it usually as the simplest explanations, which takes us to: Fungal prions en.wikipedia.org/wiki/Fungal_prions''Fungal prions provide an excellent model for the understanding of disease-forming mammalian prions. Fungal prions are naturally occurring proteins that can undergo a structural conversion that becomes self-propagating and infectious. They represent an epigenetic phenomenon in which information is not encoded in the nuclear DNA, but is structurally encoded within the protein. Several prion-forming proteins have been identified in fungi, primarily in the yeast Saccharomyces cerevisiae. Some of these are not associated with any disease state and may possibly have a beneficial role by giving an evolutionary advantage to their host[1]. Podospora anserina is a filamentous fungus. Genetically compatible colonies of this fungus can merge together and share cellular contents such as nutrients and cytoplasm. A natural system of protective "incompatibility" proteins exists to prevent promiscuous sharing between unrelated colonies. One such protein, called HET-S, adopts a prion-like form in order to function properly.'' Saccharomyces cerevisiae virus L-Aen.wikipedia.org/wiki/Saccharomyces_cerevisiae_virus_L-ASaccharomyces cerevisiae virus L-A is a double-stranded RNA (dsRNA) virus of the yeast Saccharomyces cerevisiae. This virus has a single 4.6 kb genomic segment that encodes its major coat protein, Gag (76 kDa) and a Gag-Pol fusion protein (180 kDa) formed by a -1 ribosomal frameshift. L-A can support the replication and encapsidation in separate viral particles of any of several satellite dsRNAs, called M dsRNAs, each of which encodes a secreted protein toxin (the killer toxin) and immunity to that toxin. L-A and M are transmitted from cell to cell by the cytoplasmic mixing that occurs in the process of mating. Neither is naturally released from the cell or enters cells by other mechanisms, but the high frequency of yeast mating in nature results in the wide distribution of these viruses in natural isolates. Moreover, the structural and functional similarities with dsRNA viruses of mammals has made it useful to consider these entities as viruses.  The double-stranded (ds)RNA viruses represent a diverse group of viruses that vary widely in host range (humans, animals, plants, fungi, and bacteria), genome segment number (one to twelve), and virion organization (T-number, capsid layers, or turrets). Members of this group include the rotaviruses, renowned globally as the commonest cause of gastroenteritis in young children, and bluetongue virus, an economically important pathogen of cattle and sheep. In recent years, remarkable progress has been made in determining, at atomic and subnanometeric levels, the structures of a number of key viral proteins and of the virion capsids of several dsRNA viruses, highlighting the significant parallels in the structure and replicative processes of many of these viruses. yourhealthzone.blogspot.com/This is from a post at MDR about this subject and the parasitic wasp for biocontrol we've been working on: Quote Kammy: "So, these Saccharomyces cerevisiae organisms which are used in our food processings... affect us how? What are they doing to the actin and filament production?" From post # 77 The actin nucleation-promoting activity of S. cerevisiae formins has been localized to the FH1-FH2 domains. "Saccharomyces" derives from Latinized Greek and means "sugar mold" or "sugar fungus", saccharo- being the combining form "sugar-" and myces being "fungus". Wasn't it mentioned that the fibers contain Polysaccharides? If I recall right? ok...so Saccharomyces cerevisiae formins promotes the activity of actin nucleation? .................................................. .................................................. ......... Ok, from what I understand is that fermented products such as cheese, beer or/and food with Fructose syrup for example lead to high level of sugars (glucose) in our bodies which in the other hand interferes with the production of insulin (pancreas) and other hormone producing glands and also the production of actin filaments? We have also researched that this virus is/was implicated into this bio insecticidal wasp...and that actin (a protein) is involved in viral movements... Originally Posted by Kammy To complement our studies in vivo, we study viral pathogenesis and virus-host interactions at the cell and molecular levels as well. These studies primarily are focused on the role of actin, one of the most abundant cellular proteins, in these processes. Actin appears to be involved in viral movement, nucleocapsid morphogenesis, and regulation of the switch from production of one viral phenotype to the other. Accordingly, dramatic sequential changes take place in the distribution and quantity of microfilaments throughout the course of infection which correlate with different phases of virus replication. In uninfected cells, microfilaments appear to be associated primarily with the plasma membrane. Upon infection, correlated with virus uptake and independent of early virus gene expression, thickened cables of microfilament bundles appear. Next, dependent upon early virus gene expression, microfilaments appear as ventral aggregates. Finally, and dependent on late gene expression, microfilaments appear within the nucleus where they co-localize with newly synthesized capsid protein during nucleocapsid assembly." Here is another article according sup35 and hsp104: Protein-folding Diseases and Chaperone Biology web.wi.mit.edu/lindquist/pub/ChaperoneBiol.htmlHeat-shock proteins (Hsps) are molecular chaperones that are induced when organisms are exposed to high temperatures and other stresses. These stresses cause proteins to unfold and potentially aggregate, thereby creating a protein-folding crisis in the cell. Hsp chaperones help the cell cope with this crisis by binding different types of folding intermediates and interacting with them in different ways. One, Hsp104, has a unique ability to promote the disaggregation of aggregated proteins. This biochemical function is in keeping with its biological function: Hsp104 is not required for normal growth but is required for survival under extreme stress. Homologues of Hsp104 have been found in other fungi, bacteria and plants and appear to function in a similar way. Hsp104 is a member of the important AAA family of proteins. AAA-proteins function to remodel other cellular proteins and thus affect a multitude of biological processes. Their power to remodel substrates lies in their capacity to couple substrate binding to conformational change. Hsp104’s ability to remodel protein structures also plays a critical role in protein conformation-based inheritance. In order to understand how Hsp104 functions in this capacity, we are employing a variety of biochemical and genetic approaches. -------------------------------------------------------------------- Thank you Sky for putting it all together. We're getting closer. Kat |
|
|
|
Post by skyship on Mar 28, 2010 21:42:44 GMT -5
Yes, now keep in mind the inorganic center. I will put that on here later, but,
right now we are doing well, looking at organic/biological.
This some of us have gone through this stage are are into the more advanced stages
of the intergration or should I say non integration of this construction.
HSP is causing Blooms, Werner and Rothman Thomas diseases.
So we know this in and of itself was also constructed, genes of certain
genotypes are suffering from these hsproteins.
The body is being enhanced...... some are integrating this, others are not.
There is a big picture, it involves convergence, biological/chemical/geological/
physics...........both organic and inorganic.
For what purpose we do not know, but I believe it is so the body can stand
a lot of heat or a lot of cold, because there is a cold shock protein as well, plus
a lot of magnetic counteraction. An epigenetic program, survival of the fittest,
those who can integrate or not integrate, not sure which. Selection and
adaptation of the human species. To adapt one has to integrate the changes in
the altered environment, and the changes the genes make to adapt to the
environment, however, some selection could be going on as well.
However, If we can narrow the virus down, which I have not ruled out Baculo
yet. I need to find the connections between it and the s. cerevis.
This sup35p is causing a different kind of prion problem than the cjd and the tse.
It was created.
Sup35p and baculoviruses in humans.
virus could be carried in the non mendelian vector the sup35p
Thank you for the compliment.
skyship
|
|
|
|
Post by skyship on Mar 28, 2010 23:46:50 GMT -5
Prion mutation Prions are proteins and do not contain genetic material. However, prion replication has been shown to be subject to mutation and natural selection just like other forms of replication. * Aneuploidy * Antioxidant * Budgerigar colour genetics * Homeobox * Macromutation * Muller's morphs * Mutant * Mutagenesis (process changing the genetic information) * Polyploidy * Robertsonian translocation * Signature tagged mutagenesis * Site-directed mutagenesis * TILLING (molecular biology) en.wikipedia.org/wiki/Mutation#By_impact_on_protein_sequenceSome things we can check out. skyship |
|
|
|
Post by skyship on Mar 29, 2010 13:43:08 GMT -5
Okay, ready for the bomb? I have been trying to find a way to present this, but, was not sure, I could present it in such a way as to be understood. But, here goes, a process of gradual artification, or artificial substitution for natural/organic native cell transformation, has been in the works for a long time. It has been gradual, so as not to notice its effects, however Morgellons is its effect from aggregated artificial or aggregated native cells and proteins, enzymes, mitochondria, organelles, the nucleosus itself has been altered. How was this done? It began with the The Turing protocell: There is a model. I do believe our TT mentioned this once. ========================= Synthetic Turing Protocells: vesicle self-reproduction through symmetry breaking instabilities In 1952, Alan Turing published a very influential paper in Philosophical Transactions of the Royal Society, entitled 'The Chemical Basis of Morphogenesis'. In that article, Turing proposed a possible solution to the problem of how developing systems can become heterogeneous, spatially organized entities starting from an initially homogeneous state (Turing 1952; Meinhardt 1982; Murray 1989; Lengyel & Epstein 1992). Turing showed that an appropriate compromise between local reactions and long-range communication through diffusion could generate macroscopic spatial structures....... .......The interplay of both components was described in terms of a system of partial differential equations, so-called reaction-diffusion (RD) equations, namely a set 273468si14 1.1 273468si15 1.2 Here, C1 and C2 indicate the concentrations of the two morphogens and their specific molecular interaction are described by the reaction terms Phi_i(C1, C2) with i=1, 2. These reactions could be, for example, activations, inhibitions or autocatalysis and degradation. The concentrations are spatial- and time-dependent functions, i.e. C1=C1(r, t) and C2=C2(r, t). Here r indicates the coordinates of a point r belonging to Gamma where Gamma is the spatial domain where reactions occur."....... tinyurl.com/yaycktuwww.istpace.org/Web_Final_Report/WP_1_artificial_cells_conception/modeling_artificial_cells/continuum_models_of_vesicul/synthetic_turing_protocells/index.html What comes first the structure, form or the design? Is the form random, design puts order to it? Where angmorg's forms mean something. skyship |
|
|
|
Post by aqt on Mar 29, 2010 13:49:10 GMT -5
whoa Nelly there's alot here to comprehend I satrted at the beginning with baculovirus and sceriviscae www.ncbi.nlm.nih.gov/pmc/articles/PMC310331/The method involves the replication and maintenance of the baculovirus genome in the yeast Saccharomyces cerevisiae which was accomplished by the isolation of a baculovirus recombinant containing yeast ARS and CEN sequences ensuring stable replication in yeast and a URA3 selectable marker. |
|
|
|
Post by aqt on Mar 29, 2010 13:54:12 GMT -5
|
|
|
|
Post by skyship on Mar 29, 2010 14:01:23 GMT -5
CREB is from the aplysia california, which is close to dicty. The creb is the memory gene. darn kinases keeps popping up. Abstract CREB is a transcription factor critical for long-term synaptic plasticity. Intriguingly, recent work has elucidated a role for CREB, as well as upstream CREB kinases, in the control of activity-dependent neuronal survival. Additionally, analysis of the molecular pathology of polyglutamine-repeat diseases suggest that alteration of pCREB–CBP function may underlie, at least in part, the neurodegenerative process. Taken together, these new findings support the idea that Ca2+/CREB/CBP-dependent gene regulation might be a shared mechanism critical in both long-term synaptic plasticity and neuronal survival. Author Keywords: Gene regulation; Synaptic plasticity; Neuronal survival; Calcium; CREB; CBP; CaM kinase tinyurl.com/yjnu42oURA3 encodes orotidine 5-phosphate decarboxylase (ODCase), an enzyme involved in the synthesis of pyrimidine ribonucleotides. Loss of ODCase activity leads to a lack of cell growth unless uracil or uridine is added to the media (positive selection). In contrast, if 5-FOA is added to the media ODCase can convert into the toxic compound 5-fluorouracil causing death (negative selection). Since URA3 allows for both positive and negative selection, it has been developed as a genetic marker for DNA transformations and other genetic techniques in bacteria and many fungal species. It is one of the most important genetic markers in yeast genetic modification. en.wikipedia.org/wiki/URA3skyship |
|
|
|
Post by skyship on Mar 29, 2010 14:12:35 GMT -5
R1 and R2 precursors in Turing Protocell: g1 and g2 morphogens in Turing protocell: First what is in the nucleolus?How could it have been made artificial in protocell? ibrunet.com/files/unit2/cell%20structure.pdfHave to go for the core. The nucleolus! what forms the vesicle?skyship |
|
|
|
Post by skyship on Mar 29, 2010 14:33:02 GMT -5
New technologies may help biology to deal directly with current and future crises. In their book, Microcosmos: 4 Billion Years of Evolution from our Microbial Ancestors, Margulis and Sagan (1986) describe some surprising possibilities. They feel that current man is little more than communities of bacteria, modular manifestations of the nucleated cell, and that new "artificial" life forms will emerge from symbiotic fusion of biology and technology. They see this happening along three lines: genetic biotechnology, computer robotics, and biochips: ... one day soon entire suits of genes, proteins and hormones may be dovetailed in the laboratory to create new species of microbes. As we gain a greater understanding of embryology and immunology we will surely clone cells into progressively larger and more complex organisms sure to intervene in our own evolution. As computer robotics evolve smallward to become nanotechnology, collective interactions with genetic biotechnology and natural biochips could precipitate the next evolutionary phase transition: mind/tech symbiosis. Magulis or Woese? Symbiosis ( 5 kingdoms of life) or 3 kingdoms of life? We are in symbiosis? www.quantumconsciousness.org/ultimatecomputing/index.htmlskyship |
|
|
|
Post by aqt on Mar 29, 2010 14:40:35 GMT -5
Vesicles form naturally because of the properties of lipid membranes (see micelle). Most vesicles have specialized functions depending on what materials they contain. en.wikipedia.org/wiki/Vesicle_(biology) |
|
|
|
Post by skyship on Mar 29, 2010 20:25:46 GMT -5
|
|
|
|
Post by skyship on Mar 29, 2010 20:28:14 GMT -5
|
|
|
|
Post by skyship on Mar 29, 2010 20:37:13 GMT -5
Potential pathway of V6D peptide nanotube formation. Each peptide monomer is 2 nm, and the diameter of the modeled bilayer nanotube is 50 nm. Red, hydrophilic head; blue, hydrophobic tail. Each peptide may interact with one another to form closed rings, which in turn stack on top of one another, ultimately yielding a nanotube. Three nanotubes are connected to each other through a three-way junction. This phenomenon mirrors lipid microtubule structures . www.pnas.org/content/99/8/5355/F5.expansion.htmlskyship |
|
|
|
Post by skyship on Mar 29, 2010 20:50:59 GMT -5
|
|
|
|
Post by skyship on Mar 30, 2010 12:17:27 GMT -5
When we have been so worried about using stem cells, is this not a concern as well? Protocell Research and Its Implications Perspectives in Biology and Medicine - Volume 53, Number 1, Winter 2010, pp. 136-147 The Johns Hopkins University Press Abstract: If and when protocells exist, they will exhibit three requisites for life: regeneration, replication, and evolution. Based on a recent collection of articles, this essay examines two major questions: (1) should research on development of protocells continue, and (2) what are the implications of this research for our understanding of "life."On the first question, I agree with contributors that the research should continue if there are adequate and ongoing safeguards against its misuse. On the second, I believe that creation of protocells is highly likely to escalate, rather than settle, controversies about the meaning and value of life, especially human life. muse.jhu.edu/login?uri=/journals/perspectives_in_biology_and_medicine/v053/53.1.mahowald.htmlskyship Is the protocell a nanocell? |
|
|
|
Post by skyship on Apr 3, 2010 1:55:24 GMT -5
Aneuploidy: Cell division that results in an unequal separation of genetic material creates aneuploid (from the Greek terms an, meaning "not"; eu, meaning "true"; and ploid, meaning "number") cells. Examples would be human cells with 47 or 45 chromosomes. For a diploid organism, aneuploidy is usually written algebraically as 2n+1 or 2n-1, indicating an odd number of genes. www.answers.com/topic/aneuploidy===================== * Antioxidant Space-filling model of the antioxidant metabolite glutathione. The yellow sphere is the redox-active sulfur atom that provides antioxidant activity, while the red, blue, white, and dark grey spheres represent oxygen, nitrogen, hydrogen, and carbon atoms, respectively. An antioxidant is a molecule capable of slowing or preventing the oxidation of other molecules. Oxidation is a chemical reaction that transfers electrons from a substance to an oxidizing agent. Oxidation reactions can produce free radicals, which start chain reactions that damage cells. Antioxidants terminate these chain reactions by removing free radical intermediates, and inhibit other oxidation reactions by being oxidized themselves. As a result, antioxidants are often reducing agents such as thiols, ascorbic acid or polyphenols.[1] Although oxidation reactions are crucial for life, they can also be damaging; hence, plants and animals maintain complex systems of multiple types of antioxidants, such as glutathione, vitamin C, and vitamin E as well as enzymes such as catalase, superoxide dismutase and various peroxidases. Low levels of antioxidants, or inhibition of the antioxidant enzymes, cause oxidative stress and may damage or kill cells. As oxidative stress might be an important part of many human diseases, the use of antioxidants in pharmacology is intensively studied, particularly as treatments for stroke and neurodegenerative diseases. However, it is unknown whether oxidative stress is the cause or the consequence of disease. en.wikipedia.org/wiki/Antioxidant================================== * Macromutation a mutation that has a profound effect on the resulting organism, as a change in a regulatory gene that controls the expression of many structural genes. dictionary.reference.com/browse/macromutation======================= * Muller's morphs 1946 Nobel Prize winner Hermann J. Muller (1890-1967) coined the terms amorph, hypomorph, hypermorph, antimorph and neomorph to classify mutations based on their behaviour in various genetic situations en.wikipedia.org/wiki/Muller's_morphs ============================== * * Mutagenesis (process changing the genetic information)=================== * Polyploidy Polyploidy: Introduction A disorder where the cells of the body have more than two copies of each chromosome. Terms such as triploid and tetraploid refer directly to how many chromosome sets are present in the cells. Polyploidy can cause birth defects and conditions such as Down's syndrome where each cell has 3 copies of a particular chromosome instead of one. A large percentage of spontaneous abortions are a result of polyploidy. www.wrongdiagnosis.com/p/polyploidy/intro.htm=============================== * Robertsonian translocation Font Size A A A Robertsonian translocation: A common and significant type of chromosome rearrangement that is formed by fusion of the whole long arms of two acrocentric chromosomes (chromosomes with the centromere near the very end). One in about 900 babies is born with a Robertsonian translocation making it the most common kind of chromosome rearrangement known in people. All five of the acrocentric chromosomes in people -- chromosome numbers 13, 14, 15, 21 and 22 -- have been found to engage in Robertsonian translocations. However, the formation of Robertsonian translocations was discovered by Hecht and coworkers to be highly nonrandom. Far and away the most frequent forms of Robertsonian translocations are between chromosomes 13 and 14, between 13 and 21, and between 21 and 22. In balanced form, a Robertsonian translocation takes the place of two acrocentric chromosomes and results in no problems for the person carrying it. But in unbalanced form, Robertsonian translocations produce chromosome imbalance and cause syndrome of multiple malformations and mental retardation. Robertsonian translocations between chromosomes 13 and 14 lead to the trisomy 13 (Patau) syndrome. And the Robertsonian translocations between 14 and 21 and between 21 and 22 can and do result in (trisomy 21 (Down) syndrome. Robertsonian translocations are named for the America insect geneticist W.R.B. Robertson who first described this form of translocation (in grasshoppers) in 1916 and are also known as whole-arm or centric-fusion translocations or rearrangements. www.medterms.com/script/main/art.asp?articlekey=5388just some definitions for tools ... skyship |
|
|
|
Post by skyship on Apr 3, 2010 2:47:35 GMT -5
more definitions: * Signature tagged mutagenesis www.nature.com/nrg/journal/v7/n12/fig_tab/nrg1984_F1.htmlSignature-tagged mutagenesis: barcoding mutants for genome-wide screens Piotr Mazurkiewicz , Christoph M. Tang , Charles Boone & David W. Holden Abstract DNA signature tags (molecular barcodes) facilitate functional screens by identifying mutants in mixed populations that have a reduced or increased adaptation to a particular environment. Many innovative adaptations and refinements in the technology have been described since its original use with Salmonella; they have yielded a wealth of information on a broad range of biological processes — mainly in bacteria, but also in yeast and other fungi, viruses, parasites and, most recently, in mammalian cells. By combining whole-genome microarrays and comprehensive ordered libraries of mutants, high-throughput functional screens can now be achieved on a genomic scale. www.nature.com/nrg/journal/v7/n12/glossary/nrg1984.html========================= * Site-directed mutagenesis Site-directed mutagenesis is a molecular biology technique in which a mutation is created at a defined site in a DNA molecule, usually a circular molecule known as a plasmid. In general, site-directed mutagenesis requires that the wild type gene sequence be known. This technique is also known as site-specific mutagenesis or oligonucleotide-directed mutagenesis. en.wikipedia.org/wiki/Site-directed_mutagenesis================================= Note the Oligonucleotide - directed mutagenesis: Oligonucleotide-directed mutagenesis Oligonucleotide-directed mutagenesis involves the incorporation of a mutant oligonucleotide into one strand of plasmid DNA. After DNA replication of this heteroduplex plasmid, any progeny plasmids that arise by replication of the wild-type strand will be homozygous for the wild-type allele, and any plasmids that arise by replication of the mutant strand will be homozygous for the mutant allele. The resulting population of plasmids contains both wild-type and mutant plasmids that may only differ by a single base-pair mutation in a sequence which lacks an easily assayed phenotype. Furthermore, when introduced into cells the mismatch repair system often repairs the mutated base to the complementary base in the wild-type strand before it has a chance to replicate, so the mutant plasmids are underrepresented relative to the wild-type plasmids. The trick to oligonucleotide-directed mutagenesis is to identify the desired mutants. There are many different methods of oligonucleotide-directed mutagenesis, but most are minor variations on two ways of enriching for the desired mutant plasmids: (1) methods which destroy the wild-type template strand of DNA and thus favor replication of the mutant strand, and (2) methods which simply allow the wild-type and mutant strands of DNA an equal chance of replicating. The dut ung method is an example of the first approach, and the mutS method is an example of the second approach. The dut ung method: The DNA template is obtained from plasmids or phage grown in a dut ung mutant of E. coli. The dut gene encodes dUTPase which normally degrades dUTP. An elevated concentration of dUTP accumulates in dut strains, resulting in incorporation of U in place T at some positions during DNA replication. The ung gene encodes uracil N-glycosylase which normally removes U from DNA. Thus, in the double mutant U is occasionally incorporated into DNA and this error is not repaired. Because U has the same base pairing properties and the same coding properties as T, incorporation of U into DNA in place of T is not mutagenic. A mutagenic oligonucleotide is annealed to a ssDNA template obtained from the dut ung strain. This oligonucleotide serves as a primer for in vitro DNA replication. The in vitro DNA replication reaction contains DNA polymerase, dATP, dTTP, dGTP, and dCTP but no dUTP, so no U is incorporated into the newly synthesized strand of DNA. After synthesis of the second strand of DNA is completed, the ends are covalently joined by DNA ligase. The resulting dsDNA consists of the template strand which contains U residues, and the newly synthesized strand which contains the mutant bases present in the oligonucleotide primer and does not contain any U residues. This dsDNA is then transformed into an ung+ recipient cell. The uracil N-glycosylase recognizes the U residues in the DNA, and excises the U leaving apyrimidinic (AP) sites in the template strand. Presence of AP sites makes the DNA strand biologically inactive because it cannot be replicated by DNA polymerase and the DNA is cut at the AP sites by specific endonucleases. Hence, when the dsDNA is introduced into the ung+ recipient , only the mutant strand will be replicated. www.sci.sdsu.edu/~smaloy/MicrobialGenetics/topics/in-vitro-genetics/SDM.htmlWhat is an oligonucleotide? what is it made of? An oligonucleotide (from Greek prefix oligo-, "having few, having little") is a short nucleic acid polymer, typically with twenty or fewer bases. Although they can be formed by bond cleavage of longer segments, they are now more commonly synthesized by polymerizing individual nucleotide precursors. Automated synthesizers allow the synthesis of oligonucleotides up to 160 to 200 bases. The length of the oligonucleotide is usually denoted by "mer" (from Greek meros, "part"). For example, a fragment of 25 bases would be called a 25-mer. Because oligonucleotides readily bind to their respective complementary nucleotide, they are often used as probes for detecting DNA or RNA. Examples of procedures that use oligonucleotides include DNA microarrays, Southern blots, ASO analysis, fluorescent in situ hybridization (FISH), and the synthesis of artificial genes. Oligonucleotides composed of DNA (oligodeoxyribonucleotides) are often used in the polymerase chain reaction, a procedure that can greatly amplify almost any small piece of DNA. There, the oligonucleotide is referred to as a primer, allowing DNA polymerase to extend the oligonucleotide and replicate the complementary strand. See also * Aptamer — oligonucleotides with important biological applications * Morpholino — oligos with non-natural backbones, which do not activate RNase-H but can reduce gene expression or modify RNA splicing * Polymorphism — the appearance in a population of the same gene in multiple forms because of mutations; can often be tested with ASO probes * Polynucleotide * CpG Oligodeoxynucleotide - an ODN with immunostimulatory properties en.wikipedia.org/wiki/Oligonucleotide====== Probes Primers, arrays, Direct mutations is like forced evolution, directed change or integration into dna. ==================== * TILLING (molecular biology) TILLING (Targeting Induced Local Lesions in Genomes) is a method in molecular biology that allows directed identification of mutations in a specific gene. TILLING was introduced in 2000, using the model plant Arabidopsis. TILLING has since been used as a reverse genetics method in other organisms such as zebrafish, corn, wheat, rice, soybean, tomato and lettuce. The method combines a standard technique of mutagenesis with a chemical mutagen such as Ethyl methanesulfonate (EMS) with a sensitive DNA screening-technique that identifies single base mutations (also called point mutations) in a target gene. The first paper describing TILLING used dHPLC to identify mutations (McCallum et al., 2000a) . The method was made more high throughput by using the restriction enzyme Cel-I combined with a gel based system to identify mutations (Colbert et al.,2001). Other methods of mutation detection, such as resequencing DNA, have been combined for TILLING. en.wikipedia.org/wiki/TILLING_(molecular_biology)skyship |
|
|
|
Post by skyship on Apr 4, 2010 22:08:40 GMT -5
Hello all, Sammy over on LB posted this paper, from 1900. I find it quite interesting and have read the whole thing, except for some of the cases, but, so far what I find describes the beginning of this a spore that turns into a protozoan like parasite. jem.rupress.org/cgi/reprint/6/4-6/443.pdfThe paper is a review of Rixford and Gilchrist's work by W. Ophuls, M.D. and Dr. H. C. Moffitt. The photos on the last pages are significant. prototheca and s. cerevisc. is mentioned as well, and he makes a point that some of this entirely represents s. cerev. That keyed my interest and now have reached the point of the arbuscules, that Kammy has posted. However, further looking takes me closer to the spitzenkorper, the spore or beginning spore form, a bud or a pedicle. He mentions the 'prickles" or bristles, correlating with with the "little haires" mentioned by Sir Thomas Browne. So, some very interesting Botany articles, I believe have been overlooked. I have alway thought that Morgellons involved a plant protein as well as hyphae, fungal, but also protozoan like protein, I believe Margulis called them "protoctists" somewhere between plant and microorganism. So much in this paper relates to Morgellons. I do wonder if this has been here before. Maybe created using the alchemy elements at many other times in history. The real papers have been either stolen or hidden away. Some now are starting to surface. I plan to pursue the spitzenkorper, or pedicle, or bud, or core particle, or cone that keeps this "spontaneous generating organism(s) replicating. I will try to summarize the paper, to save you all time. ============================== This paper says "life forms are protoplasmic" Tubercles are present and a mention of mycobacterium tuberculosis, and "protozoic dermatitis" Lesions are spread by lymph and blood current Pseudo diphtheria found Emyema Pericardium Miliary abscesses tubercles in spleen, kidneys, liver Ostitis of frontal bone and both tibias Abcesses and perforation through skin Inflammation of both knee joints, shoulder, elbows and wrists Fibrous adhesions Granulation tissue with tubercles Basilar tubercular meningitis Miliary tuberculosis of spleen and kidney Intestinal infection by ingestion of spore, or bud or pedicle Extensive lesions, 6 months duration. "Spherical, protoplasmic body, peripheral, double contoured, highly refractile membrane." "Vacuoles may be seen" Degenerate forms are in old lesions. There are either numerous vacuoles or the protoplasm shrinks sporulation cleaves to protoplasm inside capsule, breaks into two parts, these devel into four....... Makes me wonder if a stem cell is in there when the spores are produced. The original spore may be part stem cell?  ? Sickle-shaped Bioconcave disk, pyramidal forms are concave No Active motility of spores A tear in capsules discharges spores. The author thinks figure 8 forms with two parasites, fused where they touch like buds, both the same size and yet "there was never any direct continuity of the protoplasm" The protozoan like bodies produce lesions, which remain in the lesions. A white mouldy growth develops in pleura and spleen in some cases. Aerial hyphae becomes club shaped swellings and formation of spores at end of hypae form spores or buds. 3 or 4 or more formed at same time. He determined chlamydia spores not oidia He says, " ...these chlamydospores enter the body of susceptible animals, they develop into protozoon-like bodies with endogenous sporulation. He proved this: 2 ways 1. Animals were inoculated with pure cultures of the mould, which did not contain any protozoon-like bodies, and have developed a disease, in the lesions of which the protozoon-like bodies were found in pure culture and no mycelium. From these diseased spots the mould has, then again been obtained by cultivation. 2. Development of the different stages into one another has been directly traced under the microscope: formation of a mycelium from the protozoon-like bodies and their spores in the hanging drop: development of the chlamydospores into protozoon-like bodies in the subcutaneous abcess in Rabbit...." He brings in the paper by Dr. H. T. Ricketts "Blastomycetic (Oidiomycetic) dermatitis and its organisms" The author of this paper, however says they are not "oidia, but a "modification of them----- chlamydospores--the characteristic being that in the hyphae the protoplasm is concentrated at certain points in the form of spores, leaving between the spores segments of the original hyphae, which are empty, as shown in fig 25" He later says, "the chlamydospores formed in our fungus can develop either into a new vegetative form (development of hyphae from them when brought upon new media, see fig 31" or under favorable conditions ( in the animal body) into "Frucht-korper" with endogenous sporulation. ==================================== pause here....................................... -------------------------------- WOULD THIS NOT BE THE SPIT ZEN KORPER?       ?? -------------------------------- The author goes on to say " our fungus on account of this constant and typical endogenous sporulatin would belong to the ascomycetes" He believes in later work that genus is in between "hyphomycetes and blastomycetes" He goes on to say that "the disease ....belongs to the infectious granulomata and shows a great deal of similarity to glanders. .....As a name for the disease I should propose: Coccidioidal granuloma" he mentions the "frequent multiple infections in bone, ....osteomyelitis of some lumbar vertebrae with compression of the cord, osteomyelitis of the parietal bones with compression of the cerebrum" Sometimes healed portions of lesions show calcification. He compares lesions and histology as same with tubercle bacilli. So, this sounds so similar to Morgellons, and yet seems the patients had it worse due to organs infiltrated with this hyphae or mycelia spores and budding. So our spitzenkorper comes back ......... in another form, yet the same. What circular dna makes the fungus spontaneously keep reproducing or replicating? The title of this paper is: FURTHER OBSERVATIONS ON A PATHOGENIC MOULD FORMERLY DESCRIBED AS A PROTOZOON (COCCIDIOIDES IMMITIS, COCCIDIODES PYOGENES) skyship |
|
|
|
Post by skyship on Apr 5, 2010 1:00:53 GMT -5
Russel bodies, Lewy bodies, Collins bodies. Russell bodies: Plasmacytoma with abundant Russell bodies. H&E stain. Russell bodies are eosinophilic, large, homogenous immunoglobulin-containing inclusions usually found in a plasma cell undergoing excessive synthesis of immunoglobulin; the russell body is characteristic of the distended Endoplasmic Reticulum. This is one cell variation found in Multiple Myeloma.[1] They are named for William Russell.[2][3] en.wikipedia.org/wiki/Russell_bodies======================= Lewy body Lewy bodies are abnormal aggregates of protein that develop inside nerve cells in Parkinson's disease (PD) and some other disorders. They are identified under the microscope when histology is performed on the brain. Lewy bodies appear as spherical masses that displace other cell components. There are two morphological types: classical (brain stem) Lewy bodies and cortical Lewy bodies. A classical Lewy body is an eosinophilic cytoplasmic inclusion that consists of a dense core surrounded by a halo of 10-nm wide radiating fibrils, the primary structural component of which is alpha-synuclein. In contrast, a cortical Lewy body is less well defined and lacks the halo. Nonetheless, it is still made up of alpha-synuclein fibrils. Cortical Lewy bodies are a distinguishing feature of Dementia with Lewy bodies (DLB) and may occasionally be seen in ballooned neurons characteristic of Pick's disease and corticobasal degeneration.[1] en.wikipedia.org/wiki/Lewy_body============================== Collins bodies: A key pathological finding is the presence of neuronal inclusion bodies distributed throughout the gray matter of the cerebral cortex and in certain subcortical nuclei. These inclusions are distinct from any described previously and henceforth are identified as Collins bodies. The Collins bodies can be isolated by simple biochemical procedures and have a surprisingly simple composition; neuroserpin (a serine protease inhibitor) is their predominant component. An affinity-purified antibody against neuroserpin specifically labels the Collins bodies, confirming their chemical composition. Therefore, we propose a new disease entity—familial encephalopathy with neuroserpin inclusion bodies (FENIB). The conclusion that FENIB is a previously unrecognized neurodegenerative disease is supported by finding Collins bodies in a small kindred from Oregon with familial dementia who are unrelated to the New York family. The autosomal dominant inheritance strongly suggests that FENIB is caused by mutations in the neuroserpin gene, resulting in intracellular accumulation of the mutant protein. ajp.amjpathol.org/cgi/content/full/155/6/1901===================================== Lafora bodies: has fibrils and filaments polyglucosan bodies may be branched granular component never has limiting membrane auto recessive myoclonic epilepsy Intraneural induction liver biopsy Zebra bodies: lysosomal storage disorder mucopolysaccharidos MPS I, II, III multiple cranial nerve liver skin water soluable Microspheres bodies: Carbon Nanotube-Adsorbed Polystyrene and Poly(methyl methacrylate) Microspheres. archderm.ama-assn.org/cgi/reprint/139/1/17.pdfspherons: Bursting Dense Microspheres (Spherons) in Alzheimer's DiseaseA Review of Studies (1980-1997) on Spherons and the Pathogenesis of Alzheimer's Disease. Averback P, Morse D, Ghanbari H. Nymox Corporation, 5516 Nicholson Lane, Suite 100A, Rockville, MD 20895, USA. This paper reviews our research studies during the past 17 years on the relationship of cerebral protein dense microspheres (DMS), termed spherons, and senile plaques (SP) in the aged human brain and in AD. Initially, correlative anatomical and pathological data suggested that spherons may evolve into SP. This led to morphometric studies which strongly supported the theory. Biochemical studies were undertaken which showed that spherons could be isolated to homogeneity from brain tissue and contained the markers associated with SP. Experiments in vitro with spherons, and with inoculation of spherons into animals, reproduced SP lesion characteristics. To test the validity of using spherons for drug screening, experimental drugs were tested, a few of which are capable of blocking the formation of spheron-induced experimental SP. www.ncbi.nlm.nih.gov/pubmed/12214009www.docstoc.com/docs/479775/Some-Bodies-in-the-Brainskyship www.mdpi.com/1420-3049/14/6/2095/pdf |
|
|
|
Post by skyship on Apr 5, 2010 1:08:04 GMT -5
Proteinoid microspheres: proteinoid proteinoid microspheres Ads by Google TRANSGENOMIC Pharmacogenomics, Biomarker and Mutation Discovery Research www.pgx.transgenomic.comProtein structure Change Quantitative Label Free Protein Analysis - In Real Time! www.farfield-group.comProteomics data analysis Compare multiple protein lists and get to biology of proteomics data www.proxeon.comProtein Modification Abs Custom abs to phos, acK, cleavage, mutations, variants, Ubc, and more www.OpenBiosystems.comA protein-like substance formed by polymerization of amino acids under inorganic conditions, such as heating to over 140°C. In the 1970s it was discovered that proteinoids could also be formed by relatively mild heating (70°C) in the presence of certain inorganic catalysts (e.g. phosphoric acid). In water, proteinoids aggregate to form small round bodies called proteinoid microspheres, or ‘protocells’. These have certain attributes of living cells, such as a differentially permeable filmlike outer layer, the ability to swell and shrink due to osmotic movements of water, and the capability for budding and binary fission. It has been proposed that such microspheres could have provided a suitable vehicle for the chemical components of life to evolve a primitive form of metabolism and pave the way for the emergence of the first living cells. science.jrank.org/pages/18571/proteinoid.htmlskyship |
|
|
|
Post by skyship on Apr 5, 2010 2:15:53 GMT -5
|
|
|
|
Post by aqt on Apr 5, 2010 10:55:16 GMT -5
|
|
|
|
Post by aqt on Apr 5, 2010 11:18:59 GMT -5
|
|